PubMed API in Python#
by Avery Fernandez and Michael T. Moen
The PubMed API, part of NCBI’s Entrez Programming Utilities (E-utilities), provides programmatic access to biomedical literature from the PubMed database, enabling retrieval of bibliographic data.
This tutorial content is intended to help facilitate academic research.
Please see the following resources for more information on API usage:
Documentation
Terms
Data Reuse
NOTE: Please see access details and rate limit requests for this API in the official documentation.
These recipe examples were tested on January 27, 2026.
Setup#
The following external libraries need to be installed into your environment to run the code examples in this tutorial:
We import the libraries used in this tutorial below:
from datetime import datetime
import matplotlib.pyplot as plt
from pprint import pprint
import requests
from time import sleep
1. Retieve the Metadata of an Article by PubMed ID#
The article we are requesting has a PubMed ID (pmid) of 27933103. The retmode parameter specifies the file format, which we specify as JSON in this example.
ESUMMARY_URL = "https://eutils.ncbi.nlm.nih.gov/entrez/eutils/esummary.fcgi"
pmid = '27933103'
params = {
'db': 'pubmed',
'id': pmid,
'retmode': 'json'
}
response = requests.get(ESUMMARY_URL, params=params)
data = response.json()
# Use pprint to examine the structure of the JSON response
pprint(data["result"][pmid], depth=1)
{'articleids': [...],
'attributes': [...],
'authors': [...],
'availablefromurl': '',
'bookname': '',
'booktitle': '',
'chapter': '',
'doccontriblist': [],
'docdate': '',
'doctype': 'citation',
'edition': '',
'elocationid': '',
'epubdate': '2016 Nov 23',
'essn': '1758-2946',
'fulljournalname': 'Journal of cheminformatics',
'history': [...],
'issn': '1758-2946',
'issue': '',
'lang': [...],
'lastauthor': 'Bara JE',
'locationlabel': '',
'medium': '',
'nlmuniqueid': '101516718',
'pages': '66',
'pmcrefcount': 33,
'pubdate': '2016',
'publisherlocation': '',
'publishername': '',
'pubstatus': '258',
'pubtype': [...],
'recordstatus': 'PubMed',
'references': [],
'reportnumber': '',
'sortfirstauthor': 'Scalfani VF',
'sortpubdate': '2016/11/23 00:00',
'sorttitle': 'programmatic conversion of crystal structures into 3d printable '
'files using jmol',
'source': 'J Cheminform',
'srccontriblist': [],
'srcdate': '',
'title': 'Programmatic conversion of crystal structures into 3D printable '
'files using Jmol.',
'uid': '27933103',
'vernaculartitle': '',
'volume': '8'}
Below, we extract from specific information from the response:
# Extract the title of the article
data["result"][pmid]["title"]
'Programmatic conversion of crystal structures into 3D printable files using Jmol.'
# Use list comprehension to compile a list of the authors
authors = [author["name"] for author in data["result"][pmid]["authors"]]
authors
['Scalfani VF',
'Williams AJ',
'Tkachenko V',
'Karapetyan K',
'Pshenichnov A',
'Hanson RM',
'Liddie JM',
'Bara JE']
2. Retrieve Metadata for a List of PubMed IDs#
First, create a list of PubMed IDs:
pmids = ["34813985", "34813932", "34813684", "34813661", "34813372", "34813140", "34813072"]
Now we can go about acquiring the data from PubMed and saving the data in a dictionary, called multi_papers:
multi_papers = {}
for pmid in pmids:
params = {
'db': 'pubmed',
'id': pmid,
'retmode': 'json'
}
try:
response = requests.get(ESUMMARY_URL, params=params)
sleep(.5) # Add a delay between API calls
# Raise an error for bad responses
response.raise_for_status()
data = response.json()
multi_papers[pmid] = data["result"][pmid]
except requests.exceptions.RequestException as e:
print(f"An error occurred for ID {pmid}: {e}")
# Show how the data is a dict with PubMed IDs as keys
pprint(multi_papers, depth=1)
{'34813072': {...},
'34813140': {...},
'34813372': {...},
'34813661': {...},
'34813684': {...},
'34813932': {...},
'34813985': {...}}
# Get title for each journal with list comprehension
sources = [multi_papers[pmid]["source"] for pmid in pmids]
sources
['Cell Calcium',
'Methods',
'FEBS J',
'Dev Growth Differ',
'CRISPR J',
'Chembiochem',
'Methods Mol Biol']
3. PubMed API Calls with Requests & Parameters#
When searching through the articles, we are given a few of ways of filtering the data. A list of all the available parameters for these requests can be found in the NCBI documentation.
In this example, we use the db and term parameters:
dbis set topubmed, which is the database we are working withtermis set to our search query
ESEARCH_URL = "https://eutils.ncbi.nlm.nih.gov/entrez/eutils/esearch.fcgi"
params = {
'db': 'pubmed',
'term': 'neuroscience intervention learning',
'retmode': 'json'
}
response = requests.get(ESEARCH_URL, params=params)
data = response.json()
# Print the number of articles found by the query
data["esearchresult"]["count"]
'33729'
# Print the number of articles returned by the query
len(data["esearchresult"]["idlist"])
20
# Print the PubMed IDs of the first 5 results
data["esearchresult"]["idlist"][:5]
['41578956', '41578865', '41578071', '41577916', '41577897']
The number of returned IDs can be adjusted with the retmax paramater:
params = {
'db': 'pubmed',
'term': 'neuroscience intervention learning',
'retmode': 'json',
'retmax': 25
}
response = requests.get(ESEARCH_URL, params=params)
data = response.json()
# Note how the number of returned articles is now 25 instead of 20
len(data["esearchresult"]["idlist"])
25
The PubMed API can also be used the query to search for an author. To do so, add [au] after the name to specify it is an author.
params = {
'db': 'pubmed',
'term': 'Darwin[au]',
'retmode': 'json'
}
response = requests.get(ESEARCH_URL, params=params)
data = response.json()
# Print total number of results of the query
data["esearchresult"]["count"]
'719'
Sorting Results#
We can use the following parameters to store the data for it to be sorted in the same API call:
usehistoryis set toy, which indicates “yes”sortis set topub date, which indicates that we are sorting by publication date
params = {
'db': 'pubmed',
'term': 'Coral Reefs',
'retmode': 'json',
'usehistory': 'y',
'sort': 'pub date'
}
response = requests.get(ESEARCH_URL, params=params)
data = response.json()
# Print the PubMed IDs of the first 5 results
data["esearchresult"]["idlist"][:5]
['41506610', '41197570', '41561915', '41536161', '41344455']
# Compare to unsorted
params = {
'db': 'pubmed',
'term': 'Coral Reefs',
'retmode': 'json'
}
response = requests.get(ESEARCH_URL, params=params)
data = response.json()
# Notice that the IDs returned are different
data["esearchresult"]["idlist"][:5]
['41570066', '41568965', '41568681', '41568679', '41568056']
Searching Based on Publication Type#
We can sort by publication type by adding AND into the search:
ANDin thetermparameter indicates that both conditions must be present[pt]specifies that the filter type is publication type
More filters can be found at PubMed Help.
params = {
'db': 'pubmed',
'term': 'stem cells AND clinical trial[pt]',
'retmode': 'json'
}
response = requests.get(ESEARCH_URL, params=params)
data = response.json()
# Print number of total results
data["esearchresult"]["count"]
'6860'
4. Visualize Publication Frequencies for Different Topics#
Frequency of Topic sortpubdate Field#
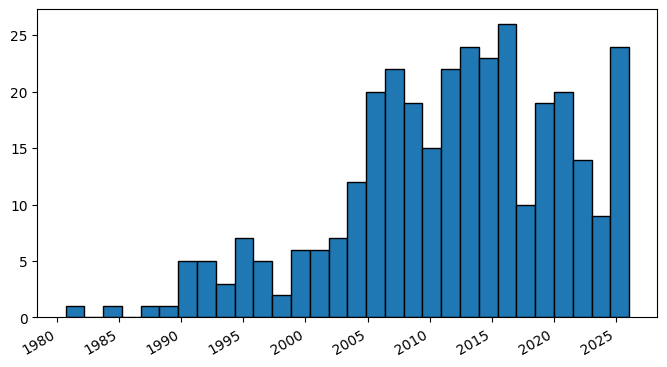
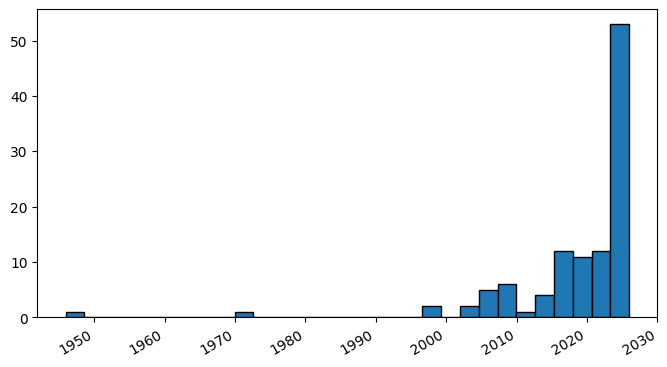
Extracting the sortpubdate field for a “hydrogel drug” search results, limited to the “clinical trial” publication type:
params = {
'db': 'pubmed',
'term': 'hydrogel drug AND clinical trial[pt]',
'retmode': 'json',
'usehistory': 'y',
'sort': 'pub date',
'retmax': 500
}
response = requests.get(ESEARCH_URL, params=params)
data = response.json()
ids = data["esearchresult"]["idlist"]
# Print the number of results
len(ids)
329
# Print the first 5 results
ids[:5]
['40730417', '41090617', '41420178', '41387884', '41168214']
# Loop through each ID and get the sortpubdate field
# Note that the sortpubdate field may not be equivalent to the publication date
pub_dates = []
for id in ids:
params = {
'db': 'pubmed',
'id': id,
'retmode': 'json'
}
try:
response = requests.get(ESUMMARY_URL, params=params)
sleep(.5)
# Raise an error for bad responses
response.raise_for_status()
data = response.json()
pub_dates.append(data["result"][id]["sortpubdate"][:10])
except requests.exceptions.RequestException as e:
print(f"An error occurred: {e}")
data = None
# Print the number of results
len(pub_dates)
329
# Print the first 5 publication dates
pub_dates[:5]
['2026/02/01', '2026/01/01', '2025/12/19', '2025/12/12', '2025/10/30']
Now that we have sortpubdates, if we want to visualize them in matplotlib, we have to convert it over to something it understands
dates = [datetime.strptime(date, '%Y/%m/%d') for date in pub_dates]
dates[:5]
[datetime.datetime(2026, 2, 1, 0, 0),
datetime.datetime(2026, 1, 1, 0, 0),
datetime.datetime(2025, 12, 19, 0, 0),
datetime.datetime(2025, 12, 12, 0, 0),
datetime.datetime(2025, 10, 30, 0, 0)]
fig, ax = plt.subplots()
plt.hist(dates, bins=30, edgecolor='black')
# set_size specifies the size of the graph
fig.set_size_inches(8, 4)
# Rotate and right-align the x labels so they don't crowd each other
for label in ax.get_xticklabels(which='major'):
label.set(rotation=30, horizontalalignment='right')
plt.show()