PubChem API in Python#
By Avery Fernandez and Michael T. Moen
PubChem provides programmatic access to chemical data and bioactivity information from the National Center for Biotechnology Information (NCBI), enabling efficient retrieval and analysis of chemical structures, identifiers, properties, and associated biological activities.
Please see the following resources for more information on API usage:
Documentation
Terms
Data Reuse
NOTE: Please see access details and rate limit requests for this API in the official documentation.
These recipe examples were tested on January 20, 2026.
Attribution: This tutorial was adapted from supporting information in:
Scalfani, V. F.; Ralph, S. C. Alshaikh, A. A.; Bara, J. E. Programmatic Compilation of Chemical Data and Literature From PubChem Using Matlab. Chemical Engineering Education, 2020, 54, 230. https://doi.org/10.18260/2-1-370.660-115508 and UA-Libraries-Research-Data-Services/MATLAB-cheminformatics
Setup#
The following external libraries need to be installed into your environment to run the code examples in this tutorial:
We import the libraries used in this tutorial below:
import pandas as pd
import requests
from time import sleep
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
1. PubChem Similarity#
Get Compound Image#
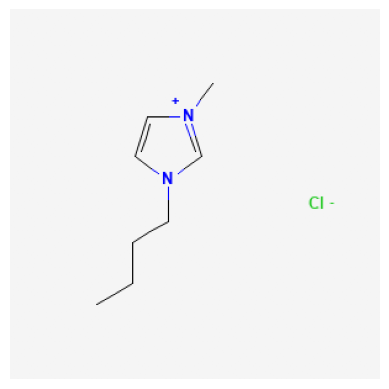
We can search for a compound and display an image. In this example, we look at 1-Butyl-3-methyl-imidazolium, which has a compound ID (CID) of 2734162.
BASE_URL = "https://pubchem.ncbi.nlm.nih.gov/rest/pug/compound/"
compoundID = "2734162"
response = requests.get(f"{BASE_URL}cid/{compoundID}/PNG")
img = response.content
# Save PNG to file
with open(f"{compoundID}.png", "wb") as out:
out.write(img)
# Display compound PNG with matplotlib
img = mpimg.imread(f"{compoundID}.png")
plt.imshow(img)
plt.axis("off")
plt.show()
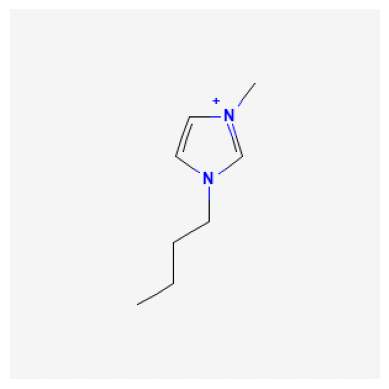
Retrieve InChI and Isomeric SMILES#
An International Chemical Identifier (InChI) is a textual representation of a substance’s molecular structure.
properties = ["IsomericSMILES", "InChI"]
response = requests.get(
f"{BASE_URL}cid/{compoundID}/property/{','.join(properties)}/JSON"
)
data = response.json()
# Display the response data
data
{'PropertyTable': {'Properties': [{'CID': 2734162,
'SMILES': 'CCCCN1C=C[N+](=C1)C',
'InChI': 'InChI=1S/C8H15N2/c1-3-4-5-10-7-6-9(2)8-10/h6-8H,3-5H2,1-2H3/q+1'}]}}
# Extract InChI
data["PropertyTable"]["Properties"][0]["InChI"]
'InChI=1S/C8H15N2/c1-3-4-5-10-7-6-9(2)8-10/h6-8H,3-5H2,1-2H3/q+1'
Isomeric SMILES is a textual representation of molecules that includes stereochemical and isotropic information.
# Extract Isomeric SMILES
data["PropertyTable"]["Properties"][0]["SMILES"]
'CCCCN1C=C[N+](=C1)C'
Perform a Similarity Search#
Search for chemical structures by similarity using a 2D Tanimoto threshold of 95% (defined by the Threshold parameter).
threshold = 95
response = requests.get(
f"{BASE_URL}fastsimilarity_2d/cid/{compoundID}/cids/JSON?Threshold={threshold}"
)
data = response.json()
id_list = data["IdentifierList"]["CID"]
# Print the number of compounds that meet the similarity threshold
len(id_list)
367
# Display first 10 results
id_list[:10]
[61347,
529334,
2734161,
118785,
12971008,
304622,
2734162,
11171745,
11424151,
11448496]
Retrieve Identifier and Property Data#
Get the following data for the retrieved compounds: InChI, IsomericSMILES, MolecularWeight, HeavyAtomCount, RotableBondCount, and Charge.
compound_data = []
properties = ["InChI", "IsomericSMILES", "MolecularWeight",
"HeavyAtomCount", "RotatableBondCount", "Charge"]
for cid in id_list[:25]:
try:
response = requests.get(
f"{BASE_URL}cid/{cid}/property/{','.join(properties)}/JSON"
)
sleep(.25)
response.raise_for_status()
data = response.json()
compound_data.append(data["PropertyTable"]["Properties"][0])
except requests.exceptions.RequestException as e:
print(f"Error fetching properties for CID {cid}: {e}")
# Convert results to a DataFrame
df = pd.DataFrame(compound_data)
df.head()
| CID | MolecularWeight | SMILES | InChI | Charge | RotatableBondCount | HeavyAtomCount | |
|---|---|---|---|---|---|---|---|
| 0 | 61347 | 124.18 | CCCCN1C=CN=C1 | InChI=1S/C7H12N2/c1-2-3-5-9-6-4-8-7-9/h4,6-7H,... | 0 | 3 | 9 |
| 1 | 529334 | 138.21 | CCCCCN1C=CN=C1 | InChI=1S/C8H14N2/c1-2-3-4-6-10-7-5-9-8-10/h5,7... | 0 | 4 | 10 |
| 2 | 2734161 | 174.67 | CCCCN1C=C[N+](=C1)C.[Cl-] | InChI=1S/C8H15N2.ClH/c1-3-4-5-10-7-6-9(2)8-10;... | 0 | 3 | 11 |
| 3 | 118785 | 110.16 | CCCN1C=CN=C1 | InChI=1S/C6H10N2/c1-2-4-8-5-3-7-6-8/h3,5-6H,2,... | 0 | 2 | 8 |
| 4 | 12971008 | 252.10 | CCCN1C=C[N+](=C1)C.[I-] | InChI=1S/C7H13N2.HI/c1-3-4-9-6-5-8(2)7-9;/h5-7... | 0 | 2 | 10 |
Retrieve Images of Compounds from Similarity Search#
# Print the first 3 result images
for cid in id_list[:3]:
try:
response = requests.get(f"{BASE_URL}cid/{cid}/PNG")
sleep(.25)
response.raise_for_status()
img = response.content
with open(f"{cid}.png", "wb") as out:
out.write(img)
except requests.exceptions.RequestException as e:
print(f"Error fetching PNG for CID {cid}: {e}")
continue
print(cid)
img = mpimg.imread(f"{cid}.png")
plt.imshow(img)
plt.axis("off")
plt.show()
61347
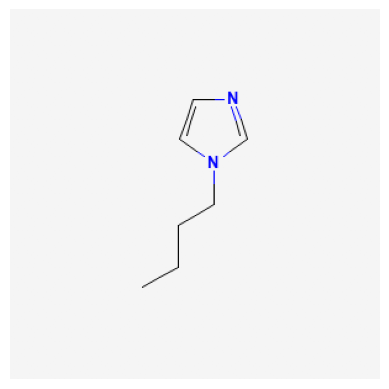
529334
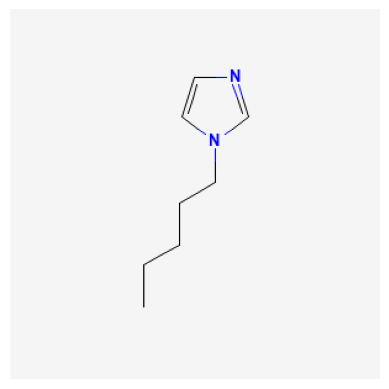
2734161
2. PubChem SMARTS Search#
We can search for chemical structures from a SMARTS substructure query using the fastsubstructure endpoint. Pattern syntax can be viewed at SMARTSPlus.
In this example, we use vinyl imidazolium substructure searches.
smartsQ = [
"[CR0H2][n+]1[cH1][cH1]n([CR0H1]=[CR0H2])[cH1]1",
"[CR0H2][n+]1[cH1][cH1]n([CR0H2][CR0H1]=[CR0H2])[cH1]1",
"[CR0H2][n+]1[cH1][cH1]n([CR0H2][CR0H2][CR0H1]=[CR0H2])[cH1]1"
]
Perform a SMARTS query search#
combinedList = []
for smarts in smartsQ:
try:
response = requests.get(f"{BASE_URL}fastsubstructure/smarts/{smarts}/cids/JSON")
sleep(.25)
response.raise_for_status()
data = response.json()
combinedList += data["IdentifierList"]["CID"]
except requests.exceptions.RequestException as e:
print(f"Error fetching substructure for SMARTS {smarts}: {e}")
request = None
# Get number of CIDs found
len(combinedList)
1024
# Print first 10 CIDs
combinedList[:10]
[2881855,
23724184,
2881236,
2881558,
2881232,
2881324,
2881449,
2881640,
24766550,
87560886]
Retrieve Identifier and Property Data#
smartsList = []
properties = ["InChI", "IsomericSMILES", "MolecularWeight",
"IUPACName", "HeavyAtomCount", "CovalentUnitCount", "Charge"]
# Demo for first 3 CIDs
for cid in combinedList[:3]:
try:
response = requests.get(f"{BASE_URL}cid/{cid}/property/{','.join(properties)}/JSON")
sleep(.25)
response.raise_for_status()
data = response.json()
smartsList.append(data["PropertyTable"]["Properties"][0])
except requests.exceptions.RequestException as e:
print(f"Error fetching properties for CID {cid}: {e}")
# Convert results to a DataFrame
df = pd.DataFrame(smartsList)
df.head()
| CID | MolecularWeight | SMILES | InChI | IUPACName | Charge | HeavyAtomCount | CovalentUnitCount | |
|---|---|---|---|---|---|---|---|---|
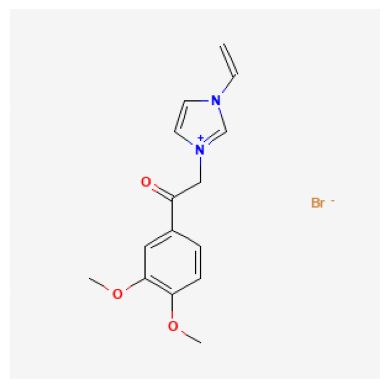
| 0 | 2881855 | 353.21 | COC1=C(C=C(C=C1)C(=O)C[N+]2=CN(C=C2)C=C)OC.[Br-] | InChI=1S/C15H17N2O3.BrH/c1-4-16-7-8-17(11-16)1... | 1-(3,4-dimethoxyphenyl)-2-(3-ethenylimidazol-1... | 0 | 21 | 2 |
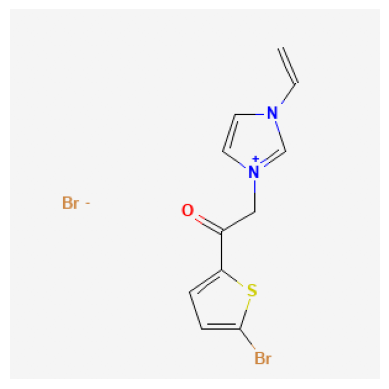
| 1 | 23724184 | 378.08 | C=CN1C=C[N+](=C1)CC(=O)C2=CC=C(S2)Br.[Br-] | InChI=1S/C11H10BrN2OS.BrH/c1-2-13-5-6-14(8-13)... | 1-(5-bromothiophen-2-yl)-2-(3-ethenylimidazol-... | 0 | 17 | 2 |
| 2 | 2881236 | 362.0 | C=CN1C=C[N+](=C1)CC(=O)C2=CC(=C(C=C2)Cl)Cl.[Br-] | InChI=1S/C13H11Cl2N2O.BrH/c1-2-16-5-6-17(9-16)... | 1-(3,4-dichlorophenyl)-2-(3-ethenylimidazol-1-... | 0 | 19 | 2 |
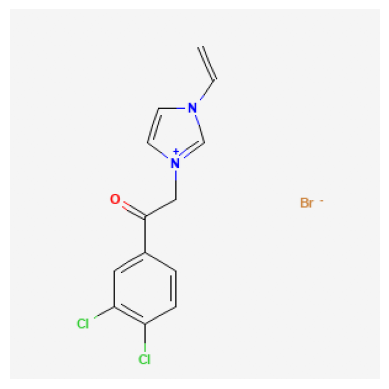
Retrieve Images of CID Compounds from SMARTS Search#
for cid in combinedList[:3]:
try:
response = requests.get(f"{BASE_URL}cid/{cid}/PNG")
sleep(.25)
response.raise_for_status()
img = response.content
with open(f"{cid}.png", "wb") as out:
out.write(img)
except requests.exceptions.RequestException as e:
print(f"Error fetching PNG for CID {cid}: {e}")
continue
# Display image
print(cid)
img = mpimg.imread(f"{cid}.png")
plt.imshow(img)
plt.axis("off")
plt.show()
2881855
23724184
2881236